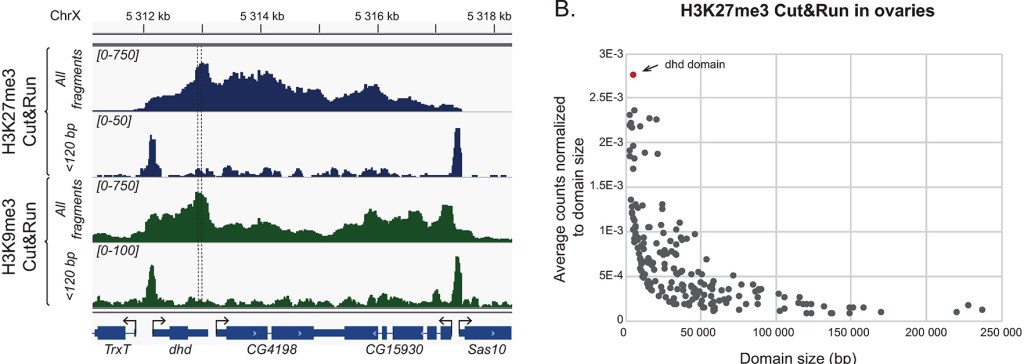
While Skene and Henikoff described this benefit of CUT&RUN, it is not easy to find a publication that takes advantage of that. But Torres-Campana et al. use it in a great paper in PLOS Genetics, 2022. A key gene in fruit fly reproduction, DHD (deadhead), is included in a very small heterochromatin domain and at least four chromatin factors participate in activation of its expression. This unique mechanism of gene activation is still incompletely understood, but precise mapping of heterochromatin marks H3K27me3 and H3K9me3 was essential and accomplished with CUT&RUN.
DHD regulation is peculiar, perhaps because it is not expressed outside ovaries and when eggs are not produced, and when expressed, it produces transcripts at a very high level. This type of need for spacial and temporal regulation is not unique, so one may expect analogous regulation in mammals.
On analytic side, Torres-Campana et al. provide novel methods required by CUT&RUN, with a different interpretation of different lengths of fragments produced in CUT step, so it is a good reading for the users of CUT&RUN. Once I understand why different lengths produce peaks in different location, I may post on that again.