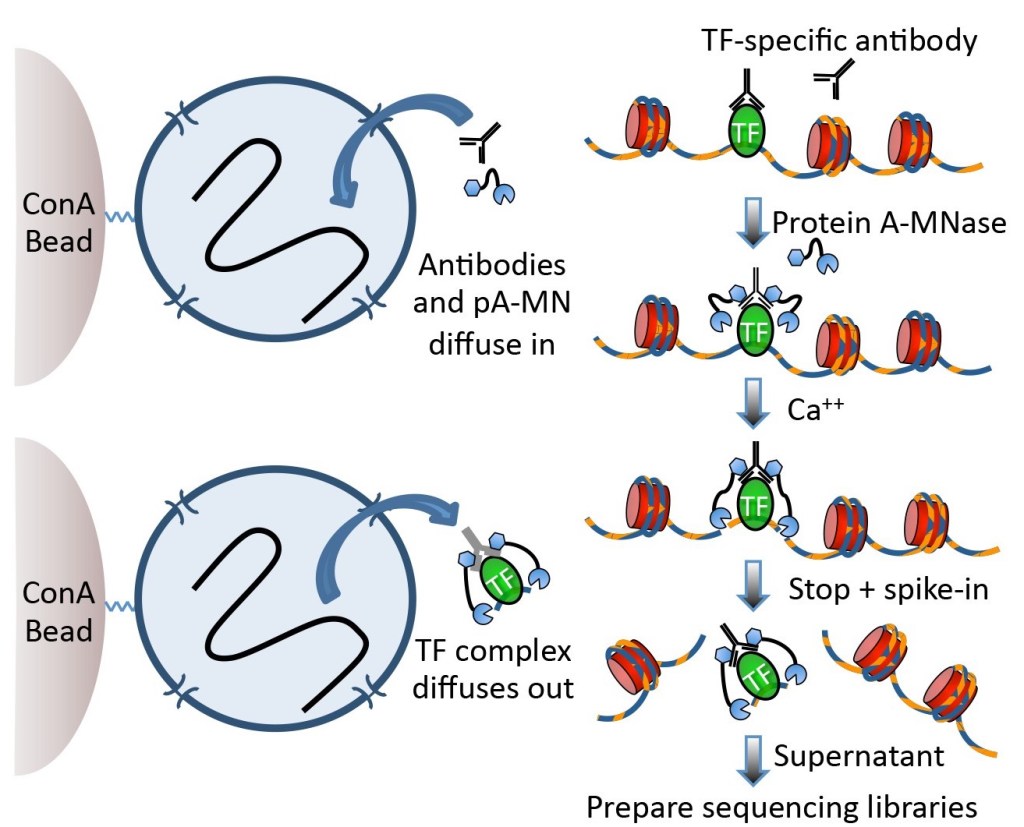
Recent papers on chromatin modification in cells increasingly use CUT&RUN of Skene and Henikoff, 2017. DNA cutting enzymes combined with an antibody enter cell nucleus and cut DNA fragments at sites where antibody binds to antigen, producing short fragments only at antigen binding sites. These short fragments exit nucleus and are collected. This yields very low off-target fraction among collected fragments and decreases the number of fragments needed to map the binding sites. Another benefit is a better mapping of binding sites in condensed chromatin which is poorly fragmented in sonication applied in ChIPseq.
While better mapping of binding sites was already used in published papers using CUT&RUN, it is less clear if it can be used for identifying 3D chromatin structure. Skene and Henikoff showed that they improved mapping of CTCF binding sites which participate in DNA-DNA contacts that define 3D structure, but it is not obvious to me how it improves the identification of pairs of such sites. Perhaps I am missing something here, or it improves the interpretation of lower resolution results obtained with existing methods.
Another intriguing possibility is to apply high-precision identification of binding sites given by CUT&RUN to predict nucleosome positions in regions marked with H3K4me3, H3K27ac etc. Nucleosome position patterns can point to changes in chromatin structure like depletion, nucleosome sliding and compacting. Perhaps this possibility was tested already.